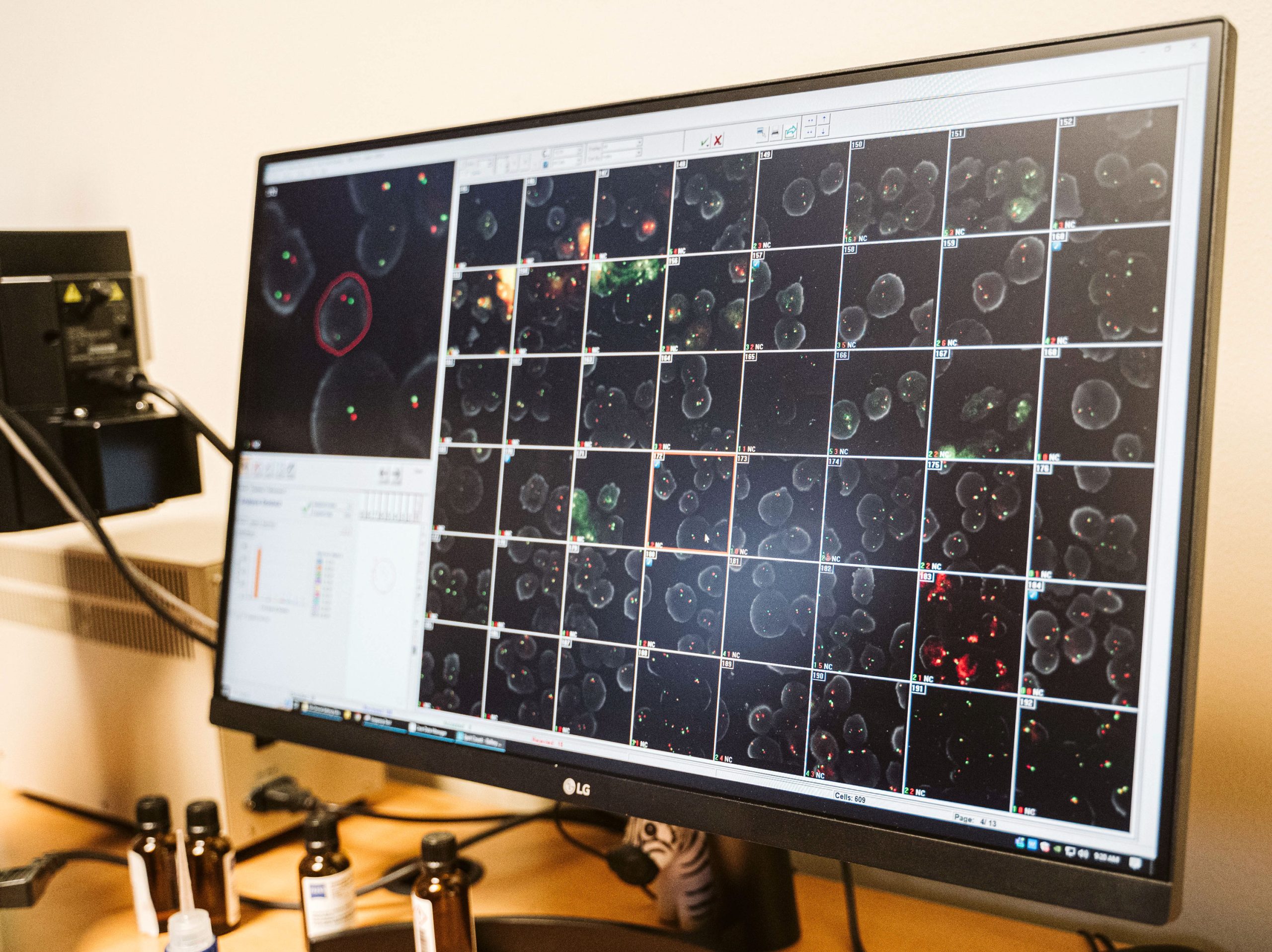
WHAT IS FLUORESCENCE IN-SITU HYBRIDIZATION (FISH)?
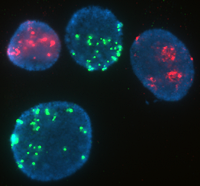 Fluorescence In-Situ hybridization (FISH) is the use of fluorochrome tagged probes that bind only to the parts of a chromosome with complementary sequences. It can be used to identify chromosome segments, to correlate chromosome structures with gene locations, to reveal cryptic abnormalities that are undetectable using standard banding techniques, and to analyze and describe complex rearrangements.
Fluorescence In-Situ hybridization (FISH) is the use of fluorochrome tagged probes that bind only to the parts of a chromosome with complementary sequences. It can be used to identify chromosome segments, to correlate chromosome structures with gene locations, to reveal cryptic abnormalities that are undetectable using standard banding techniques, and to analyze and describe complex rearrangements.
WiCell’s assays are performed by an American Board of Medical Genetics and Genomics (ABMGG) board certified or board eligible director, with results typically available in 10-15 days.
What FISH Detects:
Genomic sequence of interest
- Duplications or deletions >100kb
- 2% mosaicism (for example: cultures where >2 of 100 cells are trisomy 12)
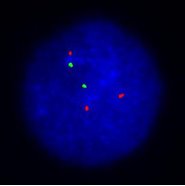
- Chromosomal location of genomic gains
- Chromosome fusions (breakaparts)
What It Doesn’t Detect:
- Changes in regions other than the probe-specific sequence
When To Use:
- To confirm findings and refine breakpoints detected by g-banded karyotyping
- To confirm findings and localize genomic gains detected by microarray analysis
- As a screen for microdeletions/duplications of known targets
- As a screen for aneuploidies
- As a screen for subtle duplications of 1q and 20q
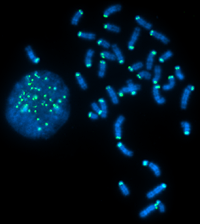
Interpreting Your Results:
When testing is complete you will receive a Fluorescence In-Situ Hybridization (FISH) Report.
cGMP and Research Grade Options Available
WiCell offers research grade and cGMP compliant FISH testing. While the assay and analysis is largely similar, the oversight, review, and systems in place to support cGMP compliant service and reporting are considerable. Stem cells intended for translational research or clinical use must meet rigorous genetic characterization release criteria for regulatory compliance. The FDA’s guidelines emphasize the importance of comprehensive cytogenetic analysis using validated methods such as Fluorescence In-Situ Hybridization analysis to ensure the safety and efficacy of stem cell products. WiCell’s cGMP FISH testing is designed to meet these exacting standards, providing the thorough documentation and quality assurance required for regulatory submissions. This level of compliance ensures that stem cell lines used in clinical trials or other translational research are thoroughly vetted for genetic integrity.
As with our non-cGMP analysis, when the cGMP FISH testing is complete, you will be provided with a cGMP FISH Analysis Report (sample normal report | sample abnormal report). As part of our cGMP FISH offering you will also receive a Certificate of Compliance which certifies your testing was completed with cGMP compliance where applicable.
If you are unsure if your project requires non-cGMP services or is ready for cGMP-grade services, this comparison chart may help you decide.